For quite some time now I’ve been tasked on multiple occasions to provide a performance analysis of some system or another. For most of these I’ve used Jupyter Notebook and now for the last couple of weeks JupyterLab without which the job at hand would have been much harder.
For those of you who haven’t yet heard of Jupyter Notebook, it’s a system that allows you to write notes in Markdown, then add some code to them (by default Python, but there are also so-called “Kernels” for other languages like Java, Go, and even Guile or Racket) which can also be used to dynamically generate diagrams and other elements that you want to show in your report.
Notebooks are made up of so-called “cells” that have a specific type. “Markdown” would be one type, while “Code” would be another. Each cell can produce output, which, in the case of code-cells, is printed below the code. Markdown cells just “overlay” the input to make the document more pleasant to read.
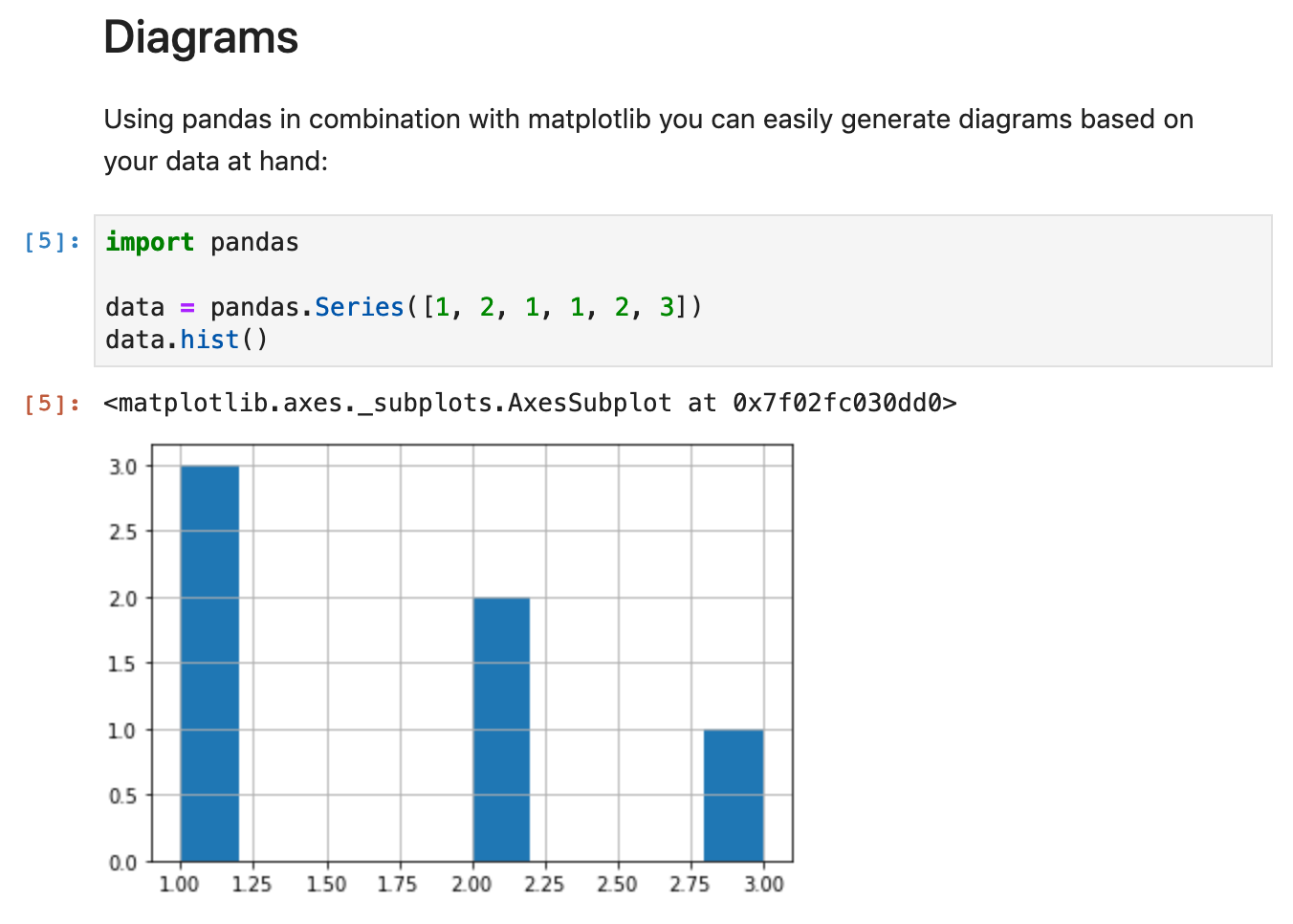
Since you can just embed Python code here, you can do pretty much anything that would make sense to be embedded inside a report (and more). For instance, I sometimes just fetch the data that should be analyzed from within the notebook using the Python Requests library.
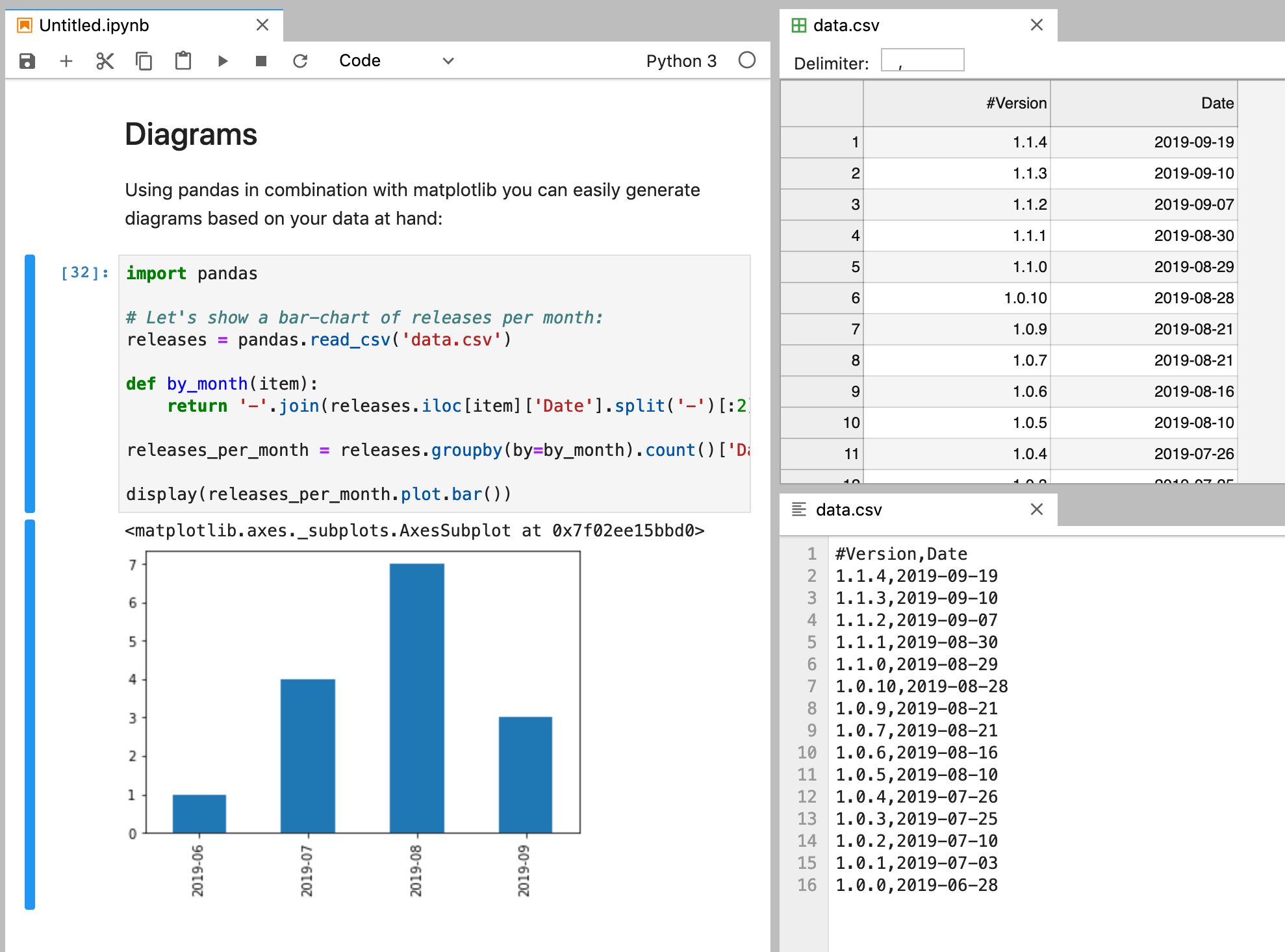
While Jupyter Notebooks feels like a simple text editor (OK, perhaps more nano than Notepad…), JupyterLab moves closer to something like VIM or Emacs (or an IDE if you prefer that kind of experience). Visually, the biggest change is that it allows you to open multiple documents side-by-side and even have cell-output being presented in a side-view.
It also comes with a preview mode for various file types, like CSV/TSV, which comes in extremely handy when you want to process such files and simply cannot keep all 40 column names in your head:
There are a couple of libraries and external binaries I use constantly, so I built myself a little utility Docker image that comes with JupyterLab and the following add-ons:
- curl
- graphviz in order to generate simple graphs using dot
- unzip
- ipywidgets
- pandas with matplotlib for powerful numerical data analysis
- folium for nice renderings of geolocations
- pendulum for more powerful datetime handling
- pyyaml in case I want to parse some YAML files as is quite common when dealing with Kubernetes environments (yes, I know that most commands also support JSON output…)
You can find this image on DockerHub under zerok/jupyter. It even comes with a little wrapper function that you can use to launch a JupyterLab instance in the current working directory:
$ eval $(docker run --rm zerok/jupyter bash-func)
$ lab
[I 17:36:49.803 LabApp] Writing notebook server cookie secret to /root/.local/share/jupyter/runtime/notebook_cookie_secret
[I 17:36:50.074 LabApp] JupyterLab extension loaded from /usr/local/lib/python3.7/site-packages/jupyterlab
[I 17:36:50.074 LabApp] JupyterLab application directory is /usr/local/share/jupyter/lab
[I 17:36:50.076 LabApp] Serving notebooks from local directory: /data
[I 17:36:50.076 LabApp] The Jupyter Notebook is running at:
[I 17:36:50.076 LabApp] http://39c8570e9c77:9980/?token=f9c906589acd04fe97130c4b34cf05c2d6950306688a9e40
[I 17:36:50.076 LabApp] or http://127.0.0.1:9980/?token=f9c906589acd04fe97130c4b34cf05c2d6950306688a9e40
[I 17:36:50.076 LabApp] Use Control-C to stop this server and shut down all kernels (twice to skip confirmation).
[C 17:36:50.079 LabApp]
I will most likely add some more libraries in the future but so far I really enjoy this setup! Perhaps this will also work for you!
During last week’s WriteTheDocs Prague I also wanted to give a little lightning talk about JupyterLab. That didn’t happen but you can still find the notebook for it on GitHub.

Do you want to give me feedback about this article in private? Please send it to comments@zerokspot.com.
Alternatively, this website also supports Webmentions. If you write a post on a blog that supports this technique, I should get notified about your link 🙂